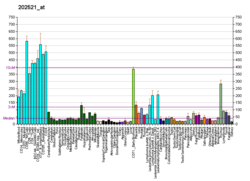
CTCF
Transcription factor
From Wikipedia, the free encyclopedia
Transcriptional repressor CTCF also known as 11-zinc finger protein or CCCTC-binding factor is a transcription factor that in humans is encoded by the CTCF gene.[5][6] CTCF is involved in many cellular processes, including transcriptional regulation, insulator activity, V(D)J recombination[7] and regulation of chromatin architecture.[8]

Discovery
Function
Although CTCF was initially discovered as a transcription factor,[9] more recent research has revealed its central role in regulating the 3D chromatin architecture.[10][11][12][8]
The role of CTCF in 3D chromatin architecture arises from its participation in the process of loop extrusion. Loop extrusion is a mechanism by which SMC complexes such as cohesin translocate along chromatin and progressively enlarge DNA loops[13][14]. In higher eukaryotes, CTCF acts as a directional barrier to extrusion: when cohesin encounters a DNA-bound CTCF oriented in the correct direction, extrusion halts. Through this mechanism, CTCF helps form chromatin loops between specific genomic elements, and establish the boundaries of topologically associating domains (TADs), thereby influencing long-range regulation of gene expression.
CTCF binding has also been both shown to promote and repress gene expression. It is unknown whether CTCF affects gene expression solely through its looping activity, or if it has some other, unknown, activity.[8] In addition to demarcating TADs, CTCF mediates promoter–enhancer interactions within a TAD.[15] It is likely that CTCF helps to bridge the transcription factors bound to transcription start sites and RNA polymerase. This model has been suggested in pioneering works on the beta-globin locus[16][8].
Evolution
CTCF is conserved across eumetazoans, including cnidarians and bilaterians, but is absent from earlier-branching animals such as sponges and from non-animal eukaryotes, suggesting that it arose in the common ancestor of eumetazoans approximately 600–700 million years ago[17].
Observed activity
The binding of CTCF has been shown to have many effects, which are enumerated below. In each case, it is unknown if CTCF directly evokes the outcome or if it does so indirectly (in particular through its looping role).
Transcriptional regulation
The protein CTCF plays a heavy role in repressing the insulin-like growth factor 2 gene, by binding to the H-19 imprinting control region (ICR) along with differentially-methylated region-1 (DMR1) and MAR3.[18][19]
Insulation
Binding of targeting sequence elements by CTCF can block the interaction between enhancers and promoters, therefore limiting the activity of enhancers to certain functional domains. Besides acting as enhancer blocking, CTCF can also act as a chromatin barrier[20] by preventing the spread of heterochromatin structures.
Regulation of chromatin architecture

CTCF plays a central role in organizing three-dimensional chromatin structure in eumetazoans by regulating the process of loop extrusion. In this mechanism, cohesin complexes translocate along chromatin and progressively enlarge DNA loops until they encounter DNA-bound CTCF. The role of CTCF as an extrusion barrier, and its function in establishing topologically associating domain (TAD) boundaries, was first proposed theoretically and later supported by experiments showing that depletion of CTCF disrupts these domain boundaries.
The barrier activity of CTCF is directional. Cohesin extrusion is halted when cohesin encounters a CTCF site in the appropriate orientation, which is determined by the orientation of the CTCF motif in the genome[21][22][23]. Structural and biochemical studies indicate that this directionality arises because the N-terminal region of CTCF interacts with cohesin and blocks its translocation along DNA.[24][25][26] Computer simulations suggests that when cohesin is halted at a CTCF site, it should continue extruding DNA from the opposite side[27]. This mechanism contributes to the formation of topologically associating domains (TADs) and other chromatin loops observed in genome-wide chromosome conformation capture experiments. Recent studies showed that such CTCF-CTCF loops are rather transient[28][29]. Stable CTCF-CTCF loops are not essential for formation of TADs, which instead arise due to continuous loop extrusion.
Through this barrier function, CTCF can both facilitate and restrict genomic interactions. CTCF sites positioned near regulatory elements can promote loop formation between them, whereas CTCF sites located between genomic regions can prevent their interaction and act as insulators[30][10][31][32]. In addition to its role as an extrusion barrier, some studies suggest that CTCF may also form homodimers[33], which may contribute to chromatin loop stabilization.
Regulation of RNA splicing
CTCF binding has been shown to influence mRNA splicing.[34]
DNA binding
CTCF binds to the consensus sequence RCCASNAGRKGGCRS (in IUPAC notation).[35][36][37] This sequence is defined by 11 zinc finger motifs in its structure. CTCF's binding is disrupted by CpG methylation of the DNA it binds to.[38] On the other hand, CTCF binding may set boundaries for the spreading of DNA methylation.[39] In recent studies, CTCF binding loss is reported to increase localized CpG methylation, which reflected another epigenetic remodeling role of CTCF in human genome.[40][41][42]
CTCF binds to an average of about 55,000 DNA sites in 19 diverse cell types (12 normal and 7 immortal) and in total 77,811 distinct binding sites across all 19 cell types.[43] CTCF's ability to bind to multiple sequences through the usage of various combinations of its zinc fingers earned it the status of a "multivalent protein".[5] More than 30,000 CTCF binding sites have been characterized.[44] The human genome contains anywhere between 15,000 and 40,000 CTCF binding sites depending on cell type, suggesting a widespread role for CTCF in gene regulation.[20][35][45] In addition CTCF binding sites act as nucleosome positioning anchors so that, when used to align various genomic signals, multiple flanking nucleosomes can be readily identified.[20][46] On the other hand, high-resolution nucleosome mapping studies have demonstrated that the differences of CTCF binding between cell types may be attributed to the differences in nucleosome locations.[47] Methylation loss at CTCF-binding site of some genes has been found to be related to human diseases, including male infertility.[36]
Protein-protein interactions
CTCF binds to itself to form homodimers.[33] CTCF has also been shown to interact with Y box binding protein 1.[48] CTCF also co-localizes with cohesin, which extrudes chromatin loops by actively translocating one or two DNA strands through its ring-shaped structure, until it meets CTCF in a proper orientation.[49] CTCF is also known to interact with chromatin remodellers such as Chd4 and Snf2h (SMARCA5).[50]